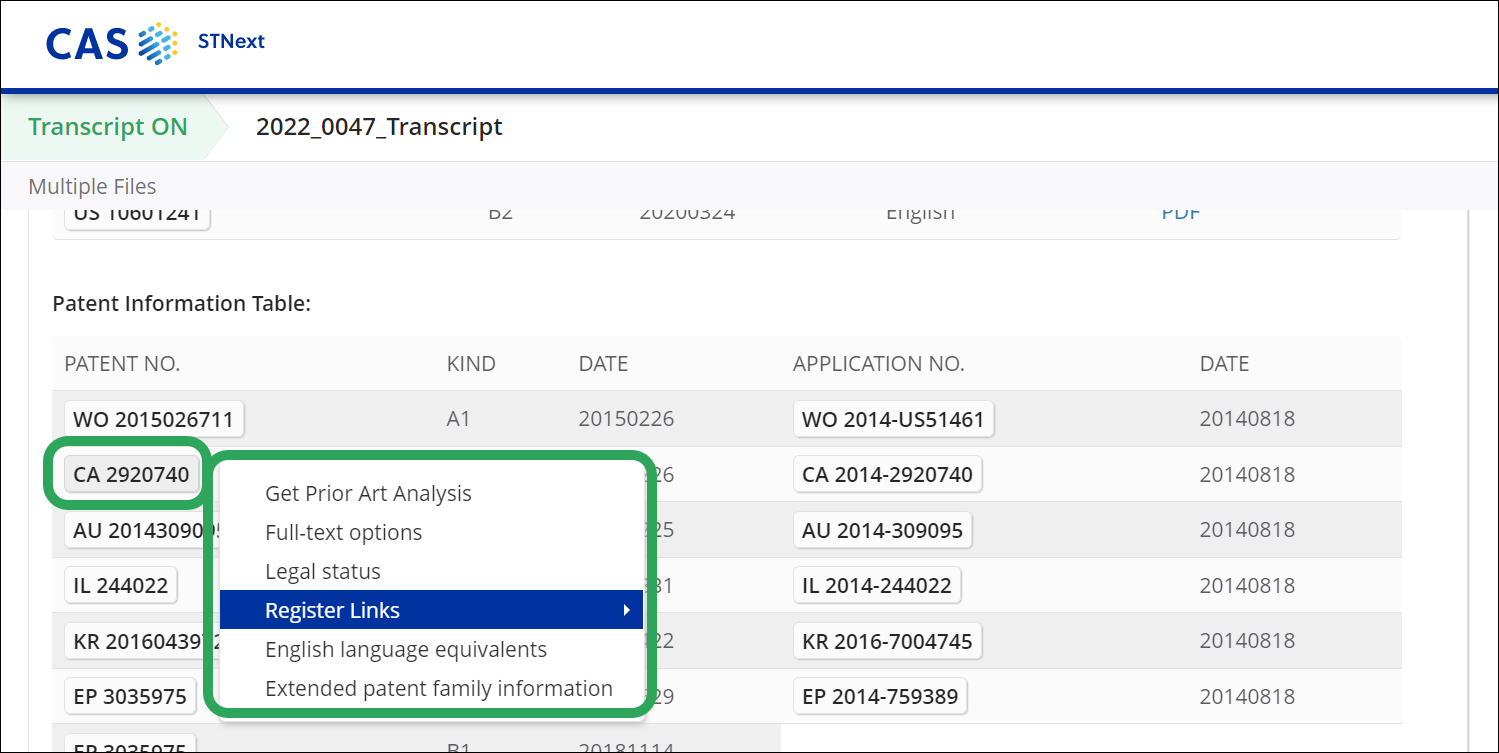
With the Register Links option, you can access direct links to register entries for national or regional patent authorities. Available in STNext databases that contain patents, the Register Links option appears in the patent number drop-down menu of publication and application numbers..

Selecting Register Links opens a submenu with a Register link.
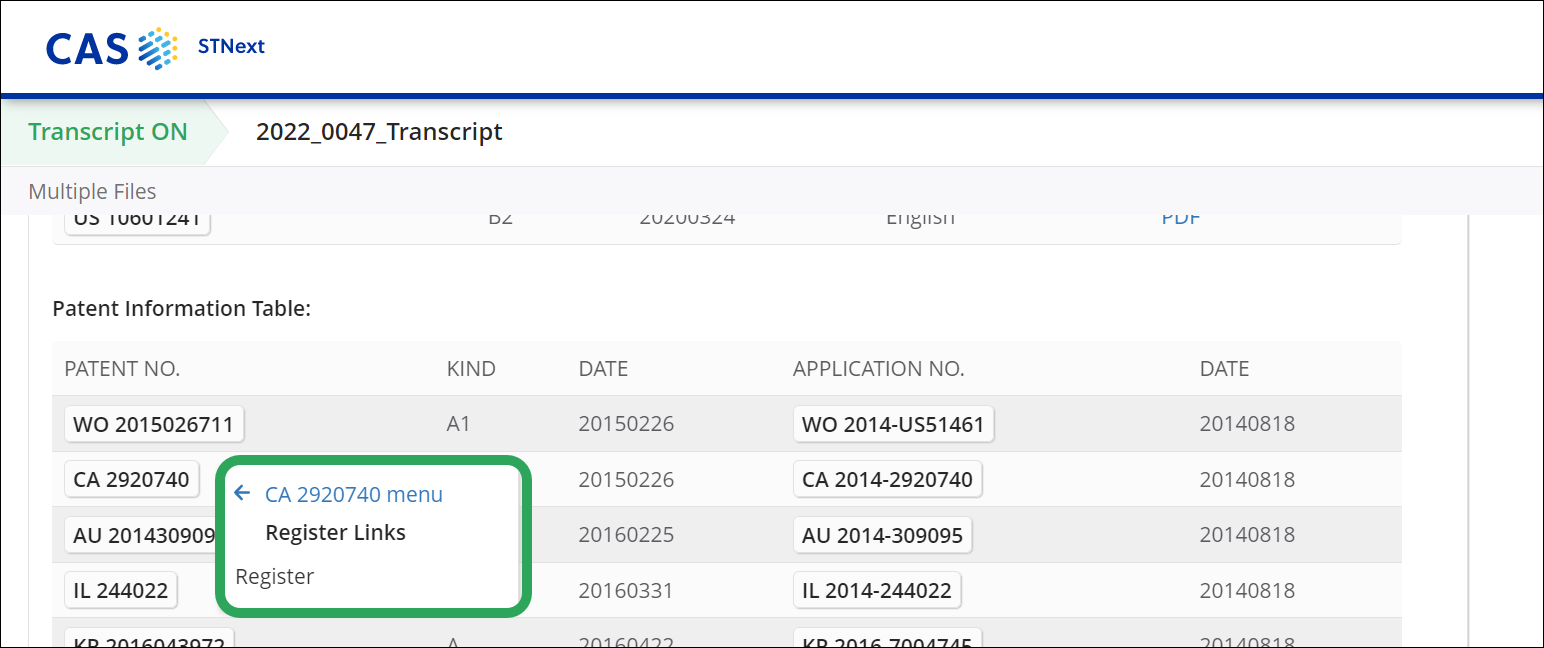
Links to patent office registers are included in about thirty databases that contain patent information. These include CAPLUS, the STN patent databases, INPADOCDB and INPAFAMDB, the DWPI databases, all STN patent full-text databases, but also all other databases that contain patent application and publication information, such as GENESEQ, USGENE or IFIALL.
The Register Links option is offered in displays that contain pairs of related application and publication numbers:
INPAFAM Patent Family table
CAPLUS Patent Information table
WPINDEX, WPIDS, WPIX Application Details
INPADOC, patent full-text databases All predefined displays such as BIB, BRIEF, ALL, MAX, and STD formats
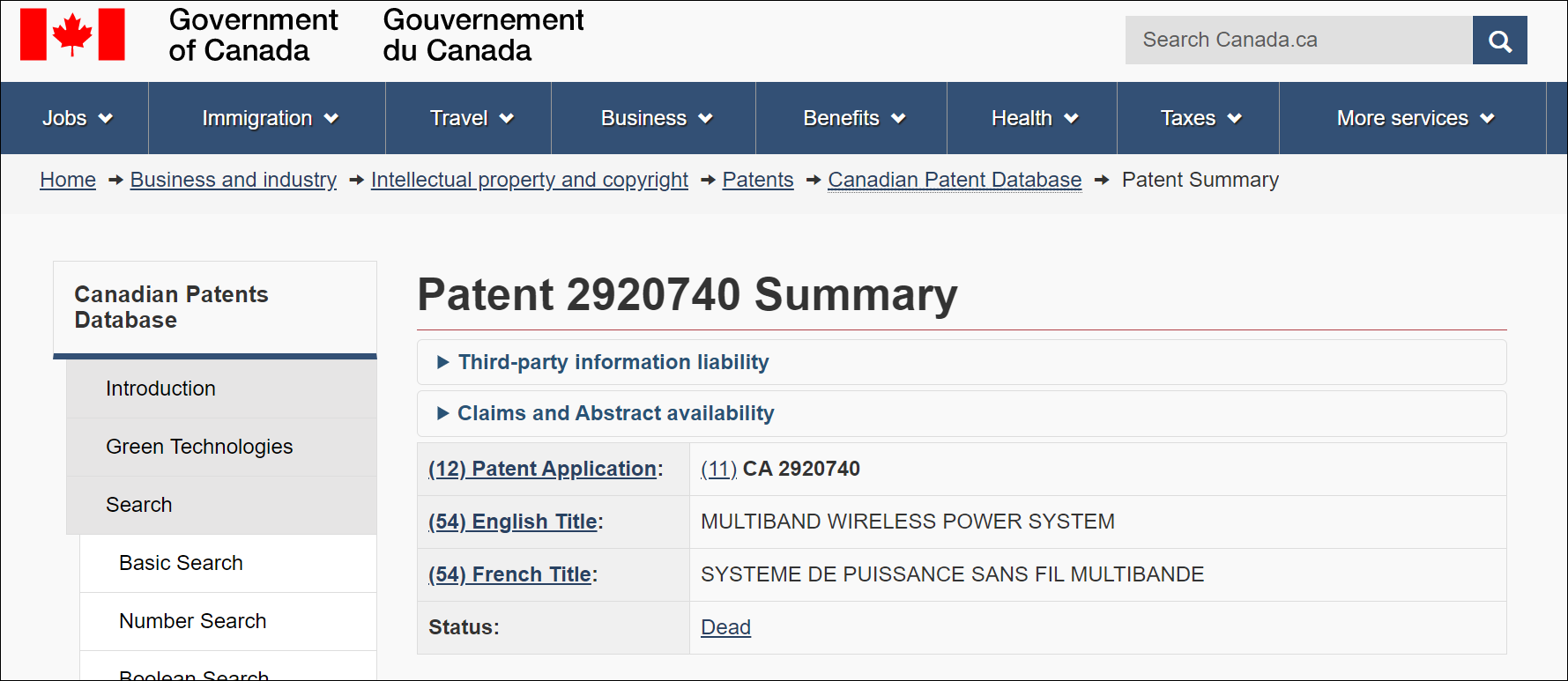
Depending on what the patent office allows, selecting Register either opens the distinct register entry of the patent application or patent office webpage in a new tab:
Linking to register entry: AT, AU, BE, CA, CH, DE, DK, EA, EE, EP, ES, FI, FR, GB, HR, IE, IL, IS, KR, LT, LU, LV, MX, MY, NL, NO, PL, SE, SK, UA, US, WO
Linking to patent
office webpage: AR, BG, CN, CZ, GR, HK, HU, IN, IT, JP, NZ,
PT, RU, SG, SI, TW
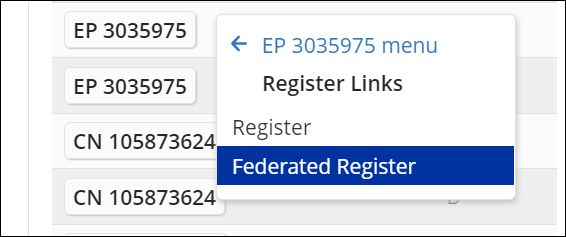
For some countries, an additional link, either to the IP5 Global Dossier (CN, JP, KR, US, and WO) or to EPO’s Federated Register is available.

When a user creates a new script via the Generate FragCode Script feature from the ellipses menu of a saved structure, FragCodes that are associated to a discontinued codes will be expanded within the Edit Script window with an OR query to include that discontinued code. For example, if code K224 is associated with the discontinued code K220, then K224 becomes (K224 OR K220) in the resulting script.
The following list shows the discontinued codes (bold) and the current codes that are associated with them:
B790: B791, B792, B793, B794, B795, B796, B797, B798
K220: K221, K222, K223, K224
K350: K351, K352, K353
K530: K531, K532, K533, K534
K600: K610, K620, K640
K800/K890: K810, K820, K830, K850
L140: L141, L142, L143, L144, L145, L146
L350: L351, L352, L353, L354, L355
L460: L461, L462, L463
M260: M261, M262, M263
M270: M271, M272, M273
M350: M351, M352, M353
M360: M361, M362, M363
M370: M371, M372, M373
M380: M381, M382, M383
Notes:
All codes except for K800, L140, L350, and L460 already appear in the FragCode scripts with their associated codes with the "OR" function.
The frag codes are only valid in the WPIX and WPIDS files.
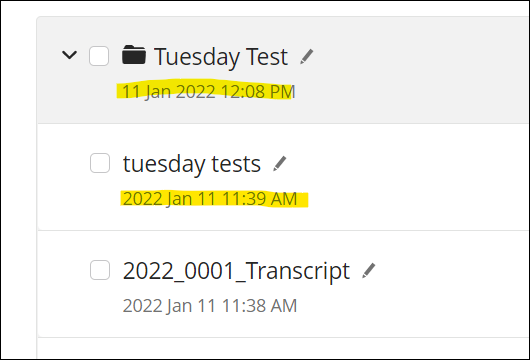
Changed My Files file dates to follow the "DD Mon YYYY" format used on folders.
Structures: 2022 Jan 11 ⇒ 11 Jan 2022
Scripts: 2022 Jan 11 ⇒ 11 Jan 2022
Biosequences: 2022 Jan 11 ⇒ 11 Jan 2022
Alerts:
January 11 2022 ⇒ 11 Jan 2022
Users can now download subject sequences in .txt or .FASTA format from sequence search results.
On the Subject tab, there is a Download Sequence button that opens a drop-down menu with TXT and FASTA options:
TXT: Entire sequence from the result downloads .txt format.
FASTA: Entire sequence from the result downloads in .FASTA format.
The file name of the download will be the name of the sequence search query with "_sequence length" directly after it where "sequence length" is the sequence length given for that result followed by the unit with no space. The possible units are either be "aa" (amino acid/proteins) or "nt" (nucleotides), (e.g., testSequence_341nt.txt).