March 11: Biosequence Results: Additional Sort Options, Maintain Filtering
and Sort Options, and Alignment Mismatch Highlighting
Additional Biosequence Result Sort Options
This release adds the new Sort By
options of Alignment Identity %,
Query Identity %,
Query Coverage %, Subject Identity
%, and Subject Coverage %.
Users may now sort a biosequence result set by:
Alignment Identity %: Ascending
Alignment Identity %: Descending
Query Coverage %: Ascending
Query Coverage %: Descending
Query Identity %: Ascending
Query Identity %: Descending
Subject Coverage %: Ascending
Subject Coverage %: Descending
Subject Identity %: Ascending
Subject Identity %: Descending

When a user selects a new sort option, the sort applies
immediately and the user returns to page one of the results if they are
not already there.
Maintain Filtering and Sorting of Biosequence Result Sets
The Filter By and Sort
By options applied to biosequence results are now retained and
will be in effect the next time the user views the result set. For Motif
search results, filter and sort options are saved with each of the child
searches.
Filter and sort options selected within Bioscape are retained when returning
to a results page.
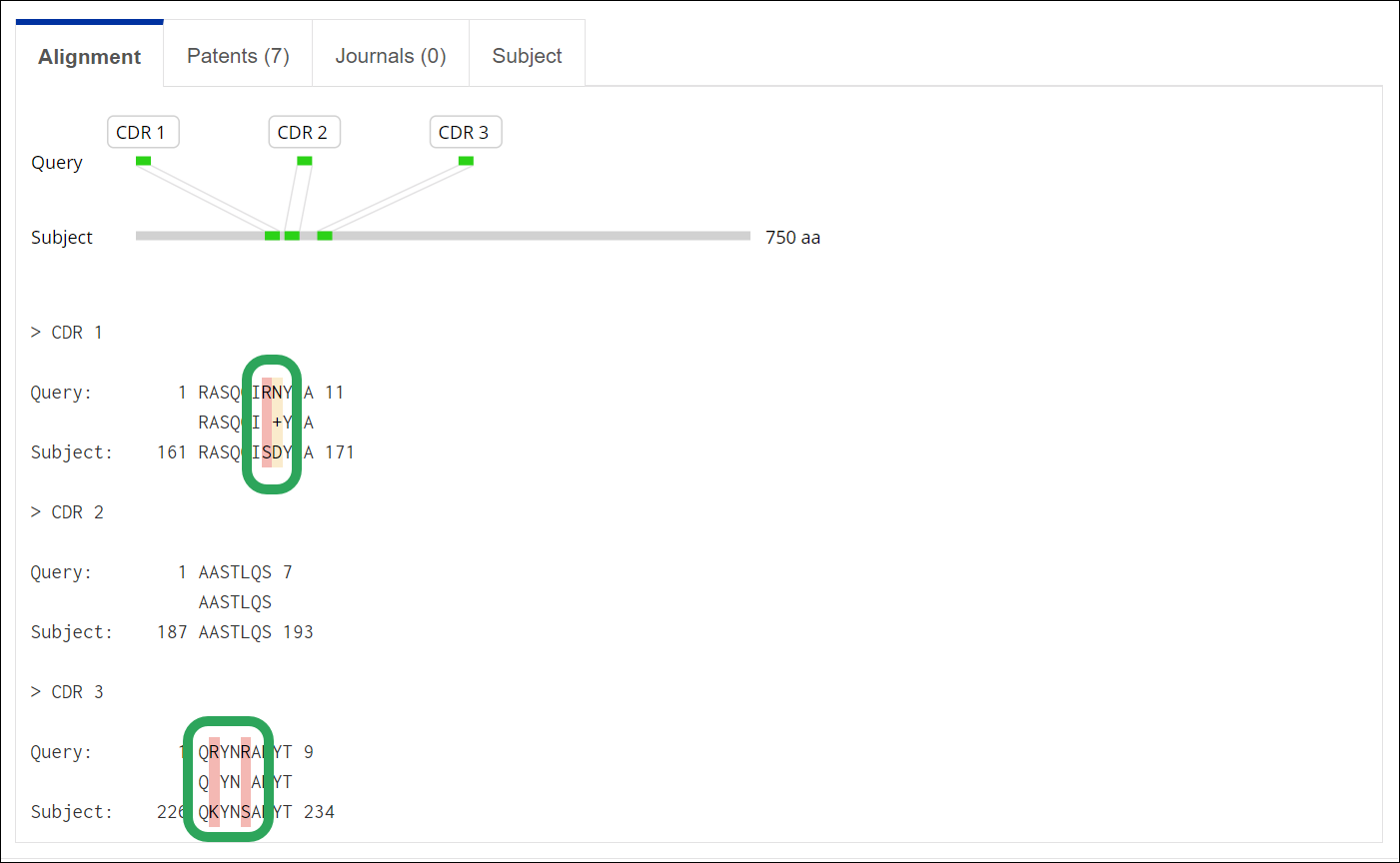
Alignment Mismatch Highlighting
Mismatches in the alignment of query (input) and subject (output) sequence
letters are now highlighted for all nucleotides and proteins searched
through the BLAST, CDR, and Motif algorithms.
Sequence mismatches between
the query and the subject are highlighted
in red; partial mismatches
are highlighted in yellow:
Mismatch highlighting in the alignment text is retained in the results
download .xlsx.

Back
to STN Application Updates